The 5th NEUBIAS Conference, held in Porto, Portugal on 8 – 12 May 2023, marked another milestone in the world of bioimage analysis and computational microscopy.
NEUBIAS is an international network that brings together researchers, bioimage analysts and developers to foster collaboration, knowledge sharing and innovation in the field of bioimage analysis. This conference, like its predecessors, aimed to explore cutting-edge techniques, novel methodologies and emerging technologies that are pushing the frontiers of bioimage analysis and beyond. The event attracted more than one hundred bioimage analysts, researchers, and experts from various academic and research institutions around the world.
The conference was organized by Paula Sampaio (i3S, PT), Rocco D’Antuono (The Francis Crick Institute, UK), Ana Stoijljkovic (BIA, CH), Aastha Mathur (Euro-Bioimaging ERIC), Beatriz Serrano Solano (Euro-Bioimaging ERIC), Maria Azevedo (i3S, PT), Kota Miura (Bioimage Analysis & Research, DE-JP), Julien Colombelli (IRB Barcelona, ES), Florian Levet (Univ. Bordeaux, FR), Elnaz Fazeli (Univ. Turku, FI), Guillaume Witz (Univ. Bern, CH), Stephane Rigaud (Institut Pasteur, FR), Hanneke Okkenhaug (Babraham Institute, UK), Bram van den Broek (Netherlands Cancer Institute, NL), and Anna Klemm (Uppsala Univ., SE).
A wide range of topics were covered, including the following:
Image Processing and Analysis: Participants discussed novel algorithms, tools, software, and cloud computing resources for efficient image processing and analysis. The integration of deep learning and artificial intelligence in bioimaging garnered significant attention, emphasizing the potential for automating complex analysis tasks.
Bioimage Data Management and Sharing: With the increasing volume of bioimaging data, discussions centred on best practices for data management, storage, and sharing to facilitate collaboration and reproducibility were welcomed.
High-Throughput and High-Content Imaging: Talks on high-throughput imaging and its applications in screening and drug discovery were particularly popular among attendees, highlighting its role in accelerating biomedical research.
Open-source Software and Tool Development: The conference provided a platform for developers to showcase their open-source bioimage analysis tools, fostering a spirit of collaboration and collective progress in the field.
The anchor of the conference was the EOSC-Life Defragmentation Training School 2, which ran in exclusivity in the first three days and shared with an open symposium in the last two, and provided an excellent opportunity for the 35 trainees to immerse in the world of cloud-based bioimage analysis solutions. Supported by the European Open Science Cloud EOSC-Life project, this training school aimed to bridge the gap between bioimage analysis and cloud computing, empowering researchers to harness the full potential of cloud technologies for their analyses. The training school featured diverse expert trainers, including bioimage analysts, cloud computing specialists, and researchers experienced in utilising cloud platforms for their work. The trainers not only shared their knowledge and expertise but also engaged in interactive discussions with the participants, addressing their queries and challenges.
The first day of the training school began with an insightful introduction to bioimage analysis by Kota Miura where he provided a comprehensive overview of the fundamental principles, multiple tools and significance of bioimage analysis in life sciences research.
 Guillaume Witz followed Kota, to introduce Jupyter notebooks and cloud-computing options. Bugra Oezdemir presented the cloud-based solutions in managing, visualizing, and sharing bioimage data efficiently using OME-Zarr format, while Peter Walczysko demonstrated the use of OMERO as a data source for cloud computing.
Guillaume Witz followed Kota, to introduce Jupyter notebooks and cloud-computing options. Bugra Oezdemir presented the cloud-based solutions in managing, visualizing, and sharing bioimage data efficiently using OME-Zarr format, while Peter Walczysko demonstrated the use of OMERO as a data source for cloud computing.
 The second day was focused on deep learning tools. It started with Marcelo Zoccoler presenting the benefits of leveraging GPUs for image processing tasks.
The second day was focused on deep learning tools. It started with Marcelo Zoccoler presenting the benefits of leveraging GPUs for image processing tasks.
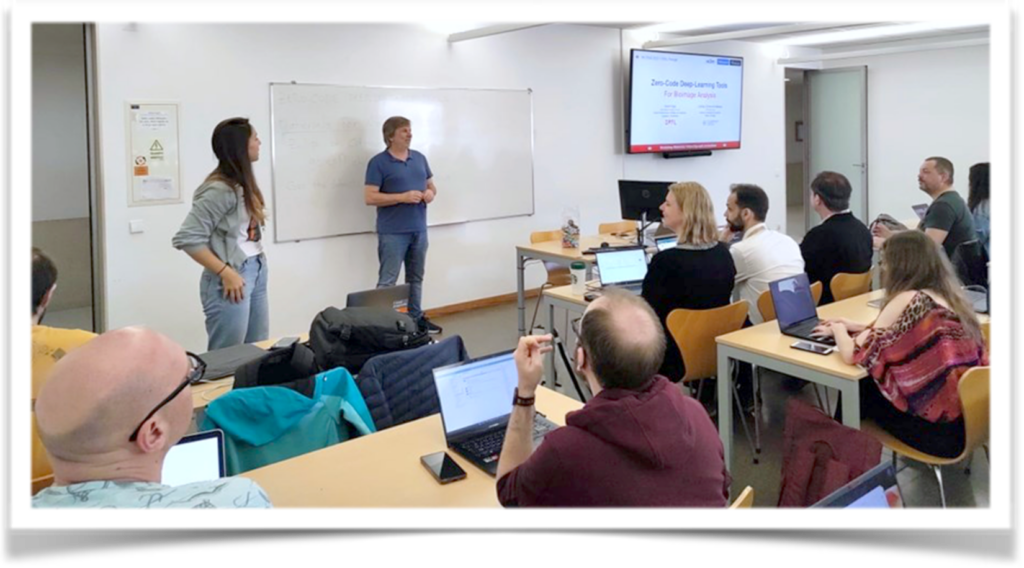
 Daniel Sage and Estibaliz Gómez de Mariscal delivered a session on “Zero Code Deep Learning Tools for Bioimage Analysis.
Daniel Sage and Estibaliz Gómez de Mariscal delivered a session on “Zero Code Deep Learning Tools for Bioimage Analysis.
 Daniel Sage also pointed out the CO2 produced by the computing resources and presented the software CodeCarbon to help track and reduce CO2 emissions from computing.
Daniel Sage also pointed out the CO2 produced by the computing resources and presented the software CodeCarbon to help track and reduce CO2 emissions from computing.
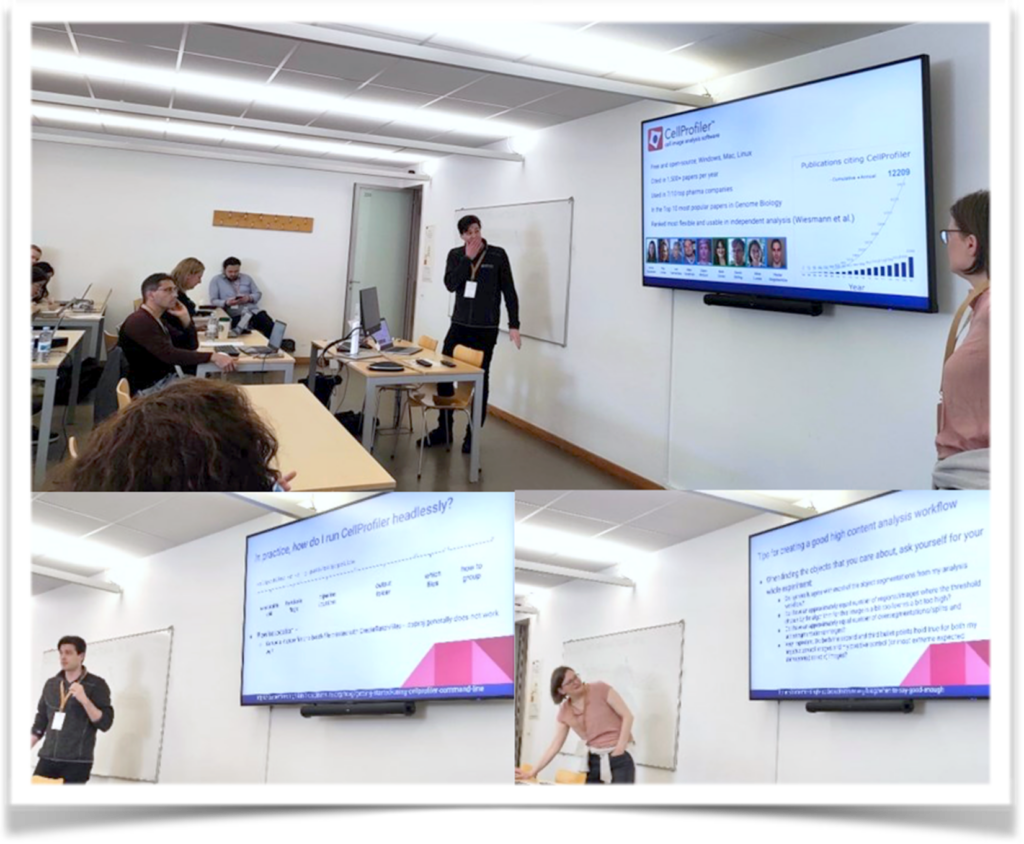
 Nodar Gogoberidze and Anna Klemm showcased the capabilities of CellProfiler software for large data analysis in a cloud environment. Also presented the Galaxy open-source platform for FAIR data analysis.
Nodar Gogoberidze and Anna Klemm showcased the capabilities of CellProfiler software for large data analysis in a cloud environment. Also presented the Galaxy open-source platform for FAIR data analysis.
 The last day of the TrainingSchool, benchmarking bioimage analysis workflows and exploring machine and deep learning on the cloud were the topics of interest. We add names such as Martin Maška speaking about the importance of benchmarking and evaluated bioimage analysis worksflow; Sebastian Tosi, Volker Beacker, and Benjamin Pavie presenting the “BIAFLOWS: A Bioimage Analysis Workflows Benchmarking Platform;
The last day of the TrainingSchool, benchmarking bioimage analysis workflows and exploring machine and deep learning on the cloud were the topics of interest. We add names such as Martin Maška speaking about the importance of benchmarking and evaluated bioimage analysis worksflow; Sebastian Tosi, Volker Beacker, and Benjamin Pavie presenting the “BIAFLOWS: A Bioimage Analysis Workflows Benchmarking Platform;
 Félix Mercier and Ignacio Arganda-Carreras conducting sessions, respectively, on Segmentation and Classification and on with clod-based machine and deep learning approaches.
Félix Mercier and Ignacio Arganda-Carreras conducting sessions, respectively, on Segmentation and Classification and on with clod-based machine and deep learning approaches.
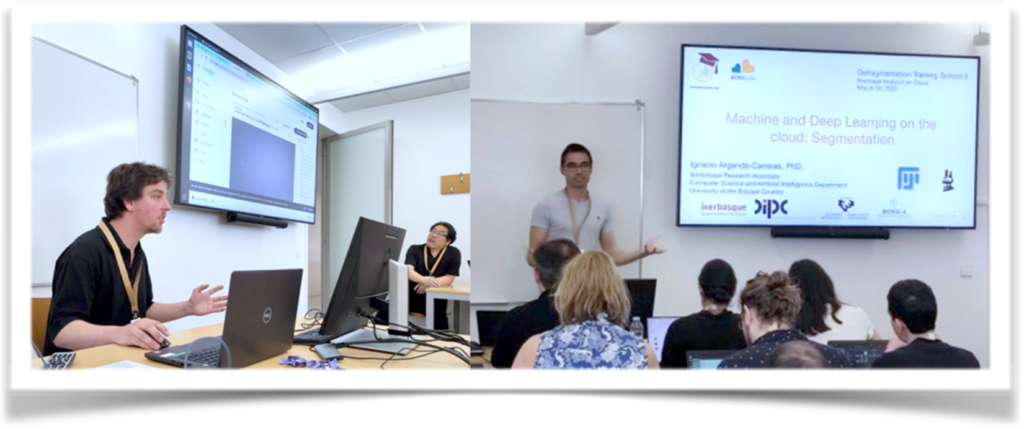 Over the three days of Training School 2, students add the opportunity to work with the different trainers on the tools they presented in the different hands-on sessions prepared. For example, students tested the OME-NGFF cloud-based workflows and publication solutions using OMERO, and they gained practical experience with “DeepImageJ”. Moreover, after the sessions, the participants worked on their own data in group settings, collaborating with trainers and helpers to address their specific bioimage analysis challenges, fostering a supportive and collaborative environment.
Over the three days of Training School 2, students add the opportunity to work with the different trainers on the tools they presented in the different hands-on sessions prepared. For example, students tested the OME-NGFF cloud-based workflows and publication solutions using OMERO, and they gained practical experience with “DeepImageJ”. Moreover, after the sessions, the participants worked on their own data in group settings, collaborating with trainers and helpers to address their specific bioimage analysis challenges, fostering a supportive and collaborative environment.


The last two days of the training school integrated the NEUBIAS Symposium which was supported as well by the AI4Live and Euro-Bioimaging. The Symposium covered a wide range of topics, including bioimage data analysis, deep learning for image data, data management, infrastructures, and high-throughput imaging data analysis. It also included two poster sessions that allowed participants to showcase their research, fostering interaction and discussion among attendees.
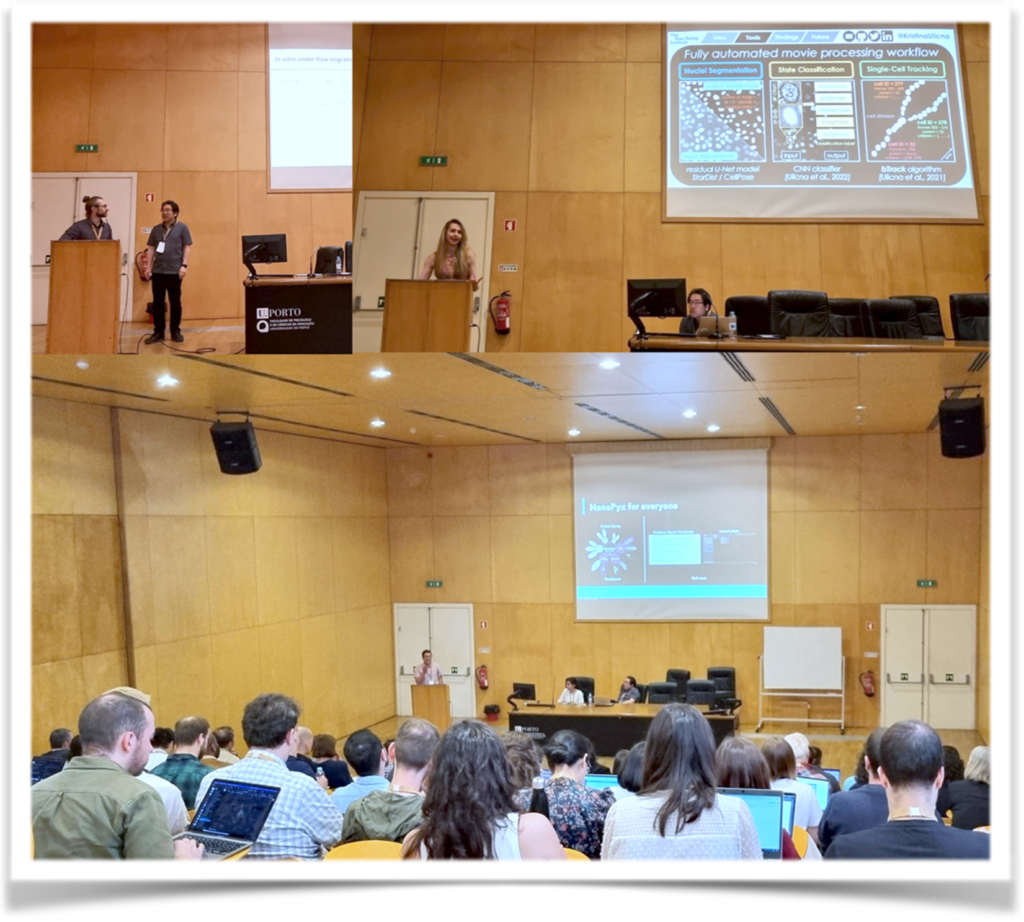
The Symposium started with Session 1 on BioImage Data Analysis, chaired by Elnaz Fazeli and Kota Miura, it brought innovative approaches in analyzing bioimaging data.Mykhailo Vladymyrov introduced UTFTrack, an advanced tool enabling the analysis of immune cell extravasation cascade across the blood-brain barrier in microfluidic devices. Kristina Ulicna presented her research on using self-supervised learning to annotate cell cycle trajectories. Stefania Marcotti shared her innovative approach to understanding tissue fibrosis using image-analysis-driven methodologies. Martin Maška provided an overview of the Cell Tracking Challenge and its significance in benchmarking cell tracking algorithms objectively. Over the past ten years, the Cell Tracking Challenge has facilitated the evaluation and comparison of various tracking methods, contributing to advancements in cell tracking accuracy and reliability. Heba Sailem introduced ShapoGraphy, a novel approach for glyph-oriented visualizing bioimaging data. Bruno M. Saraiva presented NanoPyx, a high-performance Python library designed for generating and
analysing super-resolution microscopy images.
 Florian Levet showcased PoCA, a powerful software for visualization and quantification of 3D single-molecule localization microscopy (SMLM) data. PoCA provided researchers with advanced tools to analyse SMLM datasets with high precision, facilitating a deeper understanding of molecular interactions and spatial distributions within cells. The session finished with a talk by Ross Laws that presented True 3D Analysis of Human Retinal Inner Limiting Membrane Morphology. His research utilized advanced imaging techniques to investigate the complex morphology of the retinal inner limiting membrane in three dimensions, providing new insights into retinal health and disease.
Florian Levet showcased PoCA, a powerful software for visualization and quantification of 3D single-molecule localization microscopy (SMLM) data. PoCA provided researchers with advanced tools to analyse SMLM datasets with high precision, facilitating a deeper understanding of molecular interactions and spatial distributions within cells. The session finished with a talk by Ross Laws that presented True 3D Analysis of Human Retinal Inner Limiting Membrane Morphology. His research utilized advanced imaging techniques to investigate the complex morphology of the retinal inner limiting membrane in three dimensions, providing new insights into retinal health and disease.
The NEUBIAS Symposium continued with Session 2 , supported by the AI4Life project, on Deep Learning for Image Data, chaired by Beatriz Serrano-Solano, presenting the latest advancements in deep learning techniques and their applications in bioimage analysis.
Ignacio Arganda-Carreras introduced deep learning-based bioimage analysis workflows suitable for researchers with varying levels of expertise. Estibaliz Gomez de Mariscal and Constantin Pape talks were focus on tools for easy application of deep learning algorithms to enhance cellular imaging and its analysis by researchers in the life sciences. Ko Sugawara presented his innovative work on deep learning with sparse annotations in cell image segmentation. His research demonstrated the feasibility of training accurate segmentation models with limited annotated data, enhancing the efficiency of cell image analysis. Cristina Izquierdo Lozano shared her research on data analysis for super-resolution microscopy in nanomedicine.
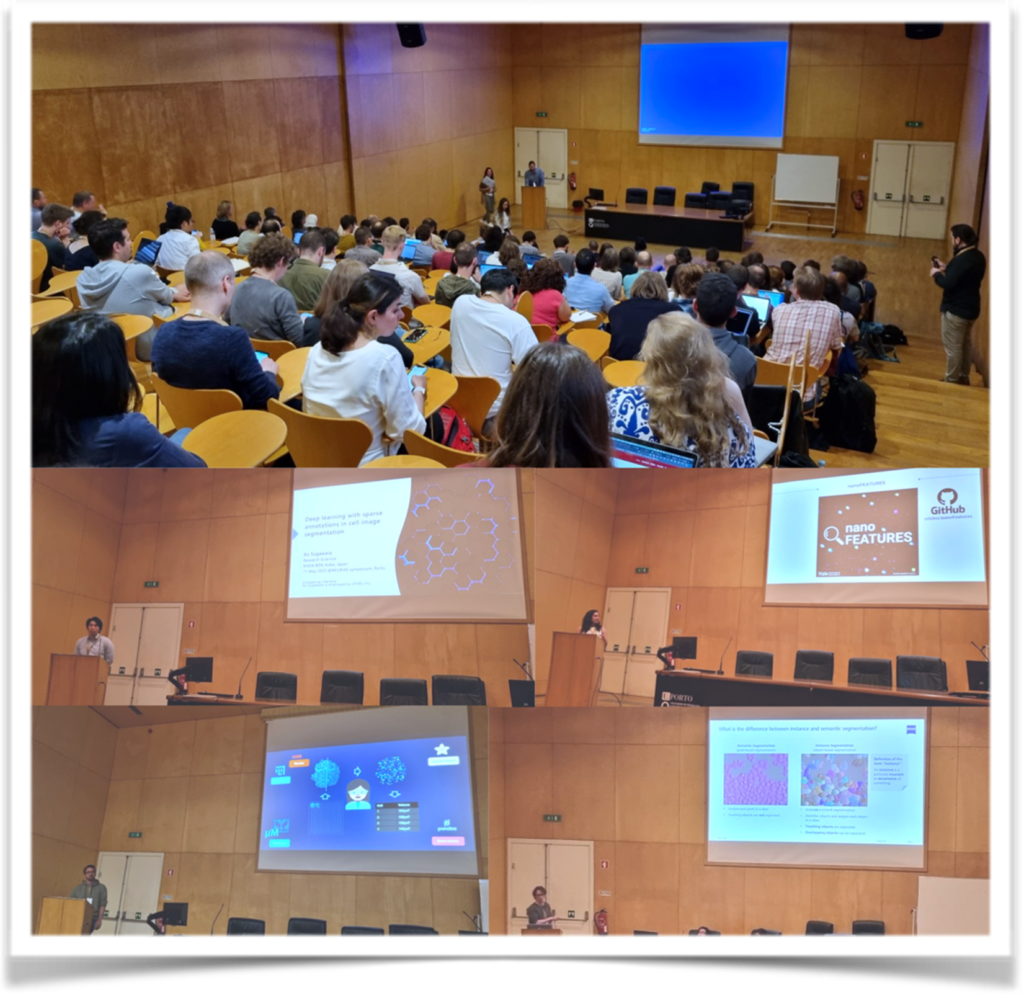 To end the session, Johannes Roos presented Arkitekt, a powerful tool for streaming analysis and real-time workflows in microscopy. His research demonstrated how Arkitekt enables researchers to efficiently process and analyze live microscopy data using deep learning methods. During the TechBites session, Sebastian Rhode presented the new AI algorithms, for instance, segmentation and a novel deployment mechanism within the ZEISS software ecosystem.
To end the session, Johannes Roos presented Arkitekt, a powerful tool for streaming analysis and real-time workflows in microscopy. His research demonstrated how Arkitekt enables researchers to efficiently process and analyze live microscopy data using deep learning methods. During the TechBites session, Sebastian Rhode presented the new AI algorithms, for instance, segmentation and a novel deployment mechanism within the ZEISS software ecosystem.
The day concluded with the Open-Source Software Lounge (OsSL), organized by Maria Azevedo and Florian Levet. The OsSL provided a platform for researchers to showcase and explore various open-source software tools and resources in the field of bioimage analysis.
Some of e OsS that were presented are the following:
- Cell-ACDC: a GUI for segmentation, tracking, cell cycle annotations and quantification of microscopy – https://github.com/SchmollerLab/Cell_ACDC
- NanoPyx: a high-performance Python library for generating and analysing super-resolution microscopy images – https://github.com/HenriquesLab/NanoPyx
- Ilastik: the interactive learning and segmentation toolkit: https://www.ilastik.org/
- BIII (Bio-Image Informatics Index): A unique database of image analysis tools and workflows for and by the bioimaging community – https://biii.eu/
- Piximi: open-source web app for performing image understanding tasks – https://www.piximi.app/
- BiaPy: Bioimage analysis pipelines in Python – https://github.com/danifranco/BiaPy
- DeepImageJ: a user-friendly plugin that enables the use of a variety of pre-trained neural networks in ImageJ and Fiji – https://deepimagej.github.io/
- Mask2Former: AI algorithm for Instance Segmentation and new deployment mechanism for AI models inside the ZEISS open software ecosystem – https://github.com/facebookresearch/Mask2Former
- Fractal: a framework to OME-Zarr process high-content imaging data at scale and prepare it for interactive visualization – https://fractal-analytics-platform.github.io/
- BIAFLOWS: A collaborative framework to benchmark BioImage Analysis workflows – https://biaflows-sandbox.neubias.org
- MoBIE: MultiModal Big Image Data Sharing and Exploration – https://mobie.github.io/
- TissUUMaps: a browser-based tool for fast visualization and exploration of millions of data points overlaying a tissue sample – https://tissuumaps.github.io/
- Webknossos: Visualization, collaborative annotation, and sharing of large 3D images – https://weblium.webknossos.org/
 Poster session and conference dinner provided a convivial atmosphere for participants to connect and build professional relationships.
Poster session and conference dinner provided a convivial atmosphere for participants to connect and build professional relationships.
 The second day of the NEUBIAS Symposium started with Session 3 focused on Data Management and Infrastructures, chaired by Aastha Mathur, with insights and experiences in managing and organizing large-scale bioimage data, fostering reproducibility, and building information infrastructures.
The second day of the NEUBIAS Symposium started with Session 3 focused on Data Management and Infrastructures, chaired by Aastha Mathur, with insights and experiences in managing and organizing large-scale bioimage data, fostering reproducibility, and building information infrastructures.
Aastha Mathur presented the efforts of Euro-BioImaging in establishing Image Data Services that adhere to the principles of Findable, Accessible, Interoperable, and Reusable (FAIR) data. Her presentation highlighted the significance of community-driven initiatives in promoting the accessibility and reproducibility of bioimage data and analysis services. Matthew Hartley discussed the BioImage Archive’s role in accelerating bioimage analysis and enhancing reproducibility. The BioImage Archive facilitates the sharing of bioimage data and metadata, enabling researchers to access and validate image datasets for their studies. Julia Fernandez-Rodriguez shared insights into the challenges and solutions related to managing correlated multimodal imaging data in life sciences. She addressed the issue of big data management for core facilities, offering strategies to handle and analyze large-scale bioimage datasets efficiently. Christian Tischer discussed the NFDI4Bioimage project, which focuses on building robust information infrastructures for imaging data.
 Beatriz Serrano-Solano introduced FAIR workflows, emphasizing the importance of creating reproducible and transparent image analysis workflows that adhere to FAIR principles. Her presentation provided valuable insights into ensuring the integrity and reliability of bioimage analysis results. Tatiana Woller highlighted the significance of FAIR data management and access to reproducible high-performance computing (HPC) environments. Adam L. Tyson presented the BrainGlobe Initiative, a project aimed at developing open-source computational neuroanatomy tools. The initiative seeks to provide researchers with powerful and accessible tools to analyze and interpret complex neuroanatomical data. The session concluded with TechBites, where Tom Herold presented on visualization, collaborative annotation, and sharing of large 3D images using WEBKNOSSOS by ScalableMinds.
Beatriz Serrano-Solano introduced FAIR workflows, emphasizing the importance of creating reproducible and transparent image analysis workflows that adhere to FAIR principles. Her presentation provided valuable insights into ensuring the integrity and reliability of bioimage analysis results. Tatiana Woller highlighted the significance of FAIR data management and access to reproducible high-performance computing (HPC) environments. Adam L. Tyson presented the BrainGlobe Initiative, a project aimed at developing open-source computational neuroanatomy tools. The initiative seeks to provide researchers with powerful and accessible tools to analyze and interpret complex neuroanatomical data. The session concluded with TechBites, where Tom Herold presented on visualization, collaborative annotation, and sharing of large 3D images using WEBKNOSSOS by ScalableMinds.
 During the event, image analysis projects based on their own research were undertaken by the seven groups of EOSC TS2 trainees. At the symposium, each group presented their achievements in bioimage analysis and highlighted the impact of the Training School on their research.
During the event, image analysis projects based on their own research were undertaken by the seven groups of EOSC TS2 trainees. At the symposium, each group presented their achievements in bioimage analysis and highlighted the impact of the Training School on their research.
 The Symposium ended with Session 4 on High-Throughput Imaging Data Analysis, chaired by Hanneke Okkenhaug and Bram van den Broek.
The Symposium ended with Session 4 on High-Throughput Imaging Data Analysis, chaired by Hanneke Okkenhaug and Bram van den Broek.
Lassi Paavolainen presented a high-content imaging-based serology test designed for SARS-CoV-2, the virus responsible for COVID-19. Bram van den Broek showcased live-cell screening of signal transduction pathways using FRET-FLIM (Fluorescence Lifetime Imaging Microscopy). Joel Luethi introduced Fractal, an open-source framework designed for processing high content imaging data stored in the OME-Zarr format. Alessia Valenti’s research focused on high-throughput image analysis to study the hormonal impact on neurodevelopment using human cortical brain organoids. Hanneke Okkenhaug presented WhiSCR, a pipeline developed for the analysis of whole-slide immunofluorescence image data using CellProfiler and R. Immanuel Sanka’s presentation focused on user-friendly droplet detection pipelines for high-throughput experiments. Hugo M. Botelho discussed high content screening from a core facility perspective. His presentation showcased the journey from experimental design to data analysis, emphasizing the crucial role of core facilities in enabling high-throughput bioimage analysis. André Maia shared insights into high content screening in a multi-thematic research environment. He presented examples and challenges faced in the analysis of high-throughput image data, emphasizing the need for versatile methodologies to address diverse research objectives. Véronique Berchet presented TechBites on cell phenotyping with Perkin Elmer Operetta® CLS™ and the Harmony high-content analysis software, showcasing advanced techniques for high-throughput cell analysis.
In conclusion, the EOSC-TS2 and NEUBIAS Symposium provided an enriching and collaborative platform for participants to explore the latest trends and advances in bioimage analysis, cloud computing, deep learning, data management and high-throughput image data analysis. The combination of taught and hands-on sessions, research talks and networking opportunities ensured a rewarding experience for all: trainees, trainers, and symposium participants. The event made a significant contribution to the advancement of bioimage analysis research and fostered collaboration between researchers, paving the way for future breakthroughs in the field. The success of the programme underlines the importance of continuous learning and knowledge exchange in the dynamic field of bioimaging.




